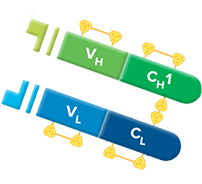
The two β-sheets pack together using the non-covalent interactions of the side chains of amino acid residues on the complementary faces. The C domains are in general compact, with short loops connecting the β-strands. The two β-sheets are covalently linked together by an intra-domain disulfide bridge formed between two cysteine residues in the ↑B and ↑F β-strands. The overall fold is often referred to as a Greek key barrel. One of the two β-sheets of the C domains has four β-strands, ↓A ↑B ↓E ↑D, and the other three β-strands, ↓C ↑F ↓G. The fold is comprised of two tightly packed anti-parallel β-sheets. The light chains are green, the heavy chains are cyan and blue, the glycan is orange sticks, and the interchain disulfides are yellow sticks.Īll the domains of heavy and light chains are approximately 110 amino acid residues in length whose conformations have been termed the “immunoglobulin fold” ( Figure 2). The engineering approaches applied to antibodies, antibody fragments, antibody, and antibody fusion products include effector function engineering, antibody humanization, affinity modulation, and stability enhancement to improve efficacy and manufacturability.Ī ribbon representation of an intact IgG, Protein Data Bank (PDB) id: 1igt, which is a mouse IgG2a isotype. Our knowledge of how antibody structure relates to function is being exploited to create antibodies and antibody-related biologics with the appropriate functional and biophysical properties to address specific therapeutic needs. The general features of antibodies described below will focus on the IgG1 framework. The other isotypes are monomeric (a monomer is defined here as a pair of HC-LCs.). The IgA and IgM isotopes have an additional J-chain, which allows the formation of dimers and pentamers, respectively. IgEs and IgMs have one variable and four constant domains. IgAs, IgDs, and IgGs have three constant (C) and one variable (V) domains. In comparison, human antibody HCs can be one of five isotypes, IgA, IgD, IgE, IgG, and IgM, each with an independent role in the adaptive immune system. Both LC classes have two domains, a constant domain (CL) and a variable domain (VL). 
Human LCs can be one of two functionally similar classes, κ or λ. The two HCs of the heterotetramer are also linked by disulfide bridges. The HC and LC of the heterodimer are linked through disulfide bonds. In natural systems, the pairing of one LC with one HC associates with another identical heterodimer to form the intact immunoglobulin. Human immunoglobulins are Y-shaped proteins composed of two identical light chains (LCs) and two identical heavy chains (HCs). The overall structure of antibodies, including the folding pattern of the individual domains and basic features of the antigen-combining sites, has been the subject of several reviews. The structural data includes complexes of these molecules with proteins, other macromolecules, peptides, and haptens. At present, the Protein Data Bank (PDB) contains over 3500 structures of antibody fragments (Fabs, Fvs, scFvs, and Fcs), as well as a small number of intact antibody structures. Our knowledge of the three-dimensional structure of antibodies has emerged from crystallographic studies reported from numerous laboratories beginning in the 1970s. The engineering approaches being used are based on our knowledge of protein structure and, in particular, our knowledge of how the structures are linked to their function. Different strategies of preparing bispecific and multispecific molecules for an array of therapeutic applications are included.Ĭurrently, all antibodies and antibody-derived macromolecules being developed for a wide spectrum of therapeutic indications require protein engineering. We also review the design and selection of binding arms, and avidity modulation. The platforms examined include the development of antibodies, antibody fragments, bispecific antibody, and antibody fusion products, whose efficacy and manufacturability can be improved via humanization, affinity modulation, and stability enhancement. In this review, our basic understanding of the antibody structure is described along with how that knowledge has leveraged the engineering of antibody and antibody-related therapeutics having the appropriate antigen affinity, effector function, and biophysical properties. Our knowledge of the structure–function relationships of antibodies provides a platform for protein engineering that has been exploited to generate a wide range of biologics for a host of therapeutic indications. Antibodies and antibody-derived macromolecules have established themselves as the mainstay in protein-based therapeutic molecules (biologics).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
